SNAREing Two Lipid Bilayers
Biological systems have a constant need for merging lipid bilayers.
This statement sounds counterintuitive at first glance: cell replication and endocytosis, the process by which white blood cells kill threatening bacteria, is the division of membranes. Why should there be value in fusing them?
The answer is surprising and ordinary. The principal example is a process that happens far more times per day than the others. The best example of our need to fuse membranes is the transmission of nerve impulses across nerve synapses. Nerve synapses are the junction boxes between two nerve cells that pass an impulse from one to the next. A good example is a simple reflex, like the connection between the thermoreceptors in our fingers and the muscles in our arms that jolt them away from the heat.
This seems a good enough reason on its own, for our bodies to have this process well worked out. However, physical chemistry is not a need-based subject: bilayers are energetically favourable, that is what drives their formation in the first place. So, what can make this process happen at the speed of a nerve impulse?
One answer is a kind of bridging protein, known as SNARE. This is one of the less tragic and more tasteful acronyms in science (compare with the organic chemists who are trying to push for diethyl azodicarboxylic acid to be referred to as DEAD). SNARE stands for Soluble NSF Attachment Protein REceptor. It is made up of several pieces (See Figure).
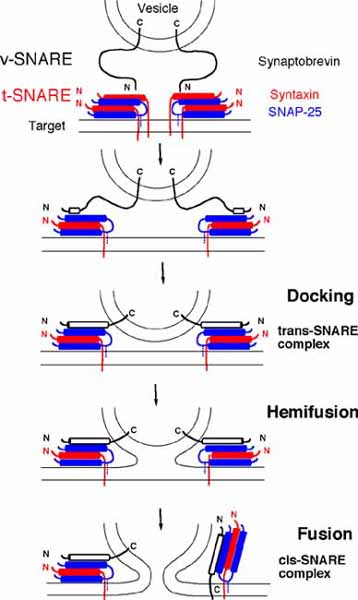
Figure. The fusion of a vesicle to a cellular membrane (labelled ‘target’ in top picture). v-SNARE, the part of the system that is attached to the vesicle at step one, interacts with t-SNARE, the part of the system on the target bilayer. The connection between these two draws the two bilayers near one another, before docking occurs. This is followed by hemi-fusion, in which the exposed monolayers of the vesicle and target begin to merge, and is followed by fusion, in which the aqueous fractions of the compartments begin to mix. Image courtesy of the University of Tennessee, http://www.bio.utk.edu/BCMB513-1/bcmb513-1.htm.
According to Thomson Reuters Web of Knowledge, the first paper in which the authors dared, and were allowed, to use this appealing acronym was published in 1994. Those authors, Sogaard et al. [1], describe a protein, called Rab, which is required for the assembly of SNARE complexes.
Other proteins, called chaperones, are required for the proper activity of SNARE [2]. This process is regulated by some of the lipids in the bilayers that are being fused: sterols, diglycerides and inositides. This implies that the membranes being fused are ‘consenting’ to being so, as without the correct mix of these lipids, the process cannot take place.
A measure of agreement by the owners of both bilayers seems fair, as the process is very quick on the scale of biological activity. Liu et al. report that it can take just 25 milliseconds [3]. That is rather less than the blink of an eye – a process that requires at least one such fusion, to occur.
What is less well understood, but has been observed in some detail is a sort of leakage that occurs during the process. It seems that the membrane properties of the two bilayers are shifted considerably with respect to their normal behaviour, as the leakage of small molecules from one or both bilayer-bound compartments during the process, can be measured [4].
We therefore know much about the process of fusing bilayers. The protein machinery, the speed at which it occurs and concurrent processes are documented. This might seem as though all of the questions have been answered and that we are but a hair’s breadth from closing the book on SNAREs. Not so: at the time of writing, the most recent work* on SNAREs to be published concerns the computational modelling of SNARE proteins, in which the molecular dynamics of a neuronal SNARE protein have been explored [5].
As well as understanding millisecond processes that we take for granted – they are of course the ones relied upon for me to type this and for you to read it– the physics of individual macromolecules can be explored as well. Perhaps only time will tell how much has yet to be enSNAREd.
References
[1] M. Segaard, K. Tani, R. R. Ye, S. Geromanos, P. Tempst, T. Kirchhausen, J. E. Rothman, T. Soellner, Cell, 1994, 78, 937.
[2] J. Mima, C. M. Hickey, H. Xu, Y.Jun, W. Wickner, EMBO J., 2008, 27, 2031.
[3] T. Liu, W. C. Tucker, A. Bhalla, E. R. Chapman, J. Weisshaar, Biophys J., 2005, 89, 2458.
[4] P. C. Zucchi and M. Zick, Mol. Biol. Cell, 2011, 22, 4635
[5] M. Tekpinar, W. Zheng, Journal of Molecular Modelling, 2014,
20, 2381.
*Retrieved 27th August, 2014.