It All Flakes off When the Lipid Shortens Tuesday, Sep 30 2014
The best word I can find for my reaction to a shoulderful of dandruff is off-putting. I have yet to meet anyone whom I know would disagree, including those who sport a pair of such shoulders themselves on a daily basis (if only they knew it). The only further thought I had had until now was an unkind one about whether or not those afflicted with this condition, use an anti-dandruff shampoo.
It is common knowledge that the white particles that we recognise as dandruff are flakes of scalp, i.e. the skin out of which our head-hair grows. The proper name for the condition is seborrheic dermatitis and it is something that affects 20% of humans of all ages. It is similar to what is called atopic dermatitis, a condition in which the non-scalp skin flakes off. It is often known as eczema. Atopic dermatitis affects 10-20% of children, but typically many fewer adults.
What I had never considered was what might cause the dandruff in the first place.
Recent research by Ishikawa et al. [1], Janssens et al. [2], Park et al. [3] and Tawada et al. [4], provide some clues. Taken together, these reports shows that the occurrence of atopic and seborrheic dermatitides correlate with a shift in the lipid profile of the skin. The evidence suggests that one lipid (with fatty acid residues that are 16 carbon atoms long, C16) is gained at the expense of one that is longer (24 carbon atoms long, C24). In other words, the hypothesis these data led to was that the fatty acids used to make these lipids are not made long enough.
In order to make sense of these observations scientifically, a recently-publsihed article by Skolova et al. [5] described the use of used a model system to research the physical impact of this change. This model was used whilst ensuring that the data generated were relevant to human skin (stratum corneum).
The model system they used consisted of the ceramide (either C16 or C24), a fatty acid (of typical chain length for mammalian lipid systems, C16) and cholesterol, also normally found in the stratum corneum. They determined the physical behaviour of the system in a comparable way using infra-red spectroscopy. This type of spectroscopy demonstrates how the fatty part of the lipid systems pack together. This technique can therefore be used to investigate the differences in packing between the C16 and C24 systems.
Both model systems behave a bit like ordinary bilayers, with the hydrophobic effect ensuring that bilayer-type systems are formed. However, the additional length of the C24 fatty acid residue in the ceramide may lead to several significant differences in the physical properties and parameters of the system. For example, the bilayer may thicken. Alternatively, the longer chain of the fatty acid residue might bend or even fold, to fill in gaps in curved systems.
In this system, which is typically rather drier than most naturally-occurring lipid systems, the effect is somewhat different. The evidence suggests that instead of a classical bilayer, the ceramide behaves more like a wax, in which the carbon chains are 180 degrees apart, rather than parallel (blue molecule, Figure). This means that the head group of the ceramide represents the hydrophilic part of the system—that which would normally be two faces of head groups and an aqueous fraction—and the system is held together as much by the van der Waal’s forces between hydrophobic moieties as it is by the hydrogen bonding and electrostatic (charge-based) forces.
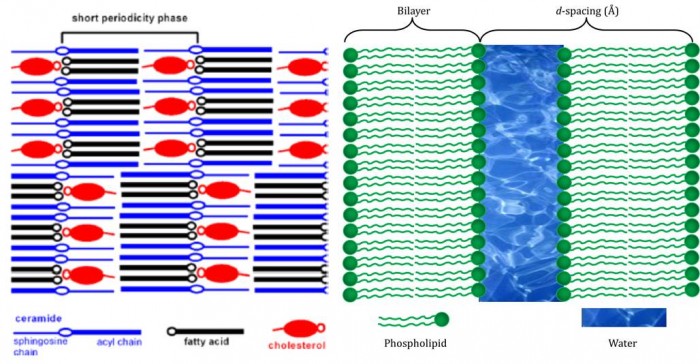
Figure. Left: diagram of the molecular assembly of ceramides, fatty acids and cholesterol in human skin. The blue molecules are not packed as classical lipids, but splayed at 180°, leaving the head group (blue ‘O’)in the middle, rather than on one face. Note how the hydrophilic parts of the cholesterol (red ‘o’) and fatty acid (black ‘o’) are assembled facing one another adjacent to that of the ceramide, forming an hydrophobic surface. (this diagram is from Skolova et al.[5], from data published by Iwai et al. [6]. Right: An ordinary bilayer. The structural parameter marked as ‘d-spacing is used to record the size and shape of systems.
It is therefore straightforward to realise how a much shorter chain—sorter by a third—might interfere with the balance of forces present in the system. We therefore expect that the skin that flakes to produce dandruff is rather weaker than normal and is the result of a faulty enzyme. The enzymes that produce longer-than-average-length fatty acids are called elongases.
Despite this insight, and how it has changed my view of what dandruff is, I must confess I cannot find use for it in conversation. Would anyone be able to say to someone with eczema “I’m sorry to see your elongases are not working at the moment”?
References
[1] J. Ishikawa, H. Narita, N. Kondo, M. Hotta, Y. Takagi, Y. Masukawa, T. Kitahara, Y. Takema, S. Koyano, S. Yamazaki, A. Hatamochi. Journal of Investigative Dermatology, 2010, 130, 2511−2514.
[2] M. Janssens, J. van Smeden, G. S. Gooris, W. Bras, G. Portale, P. J. Caspers, R. J. Vreeken, T. Hankemeier, S. Kezic, R. Wolterbeek, A. P. Lavrijsen , J. A. Bouwstra. Journal of Lipid Research, 2012, 53, 2755−2766.
[3] Y. H. Park, W. H. Jang, J. A. Seo, M. Park, T. R. Lee, Y. H.Park, D. K. Kim, K. M. Lim. Journal of Investigative Dermatology, 2012, 132, 476−479.
[4] C. Tawada, H. Kanoh, M. Nakamura, Y. Mizutani, T. Fujisawa, Y. Banno, M. Seishima/ Journal of Investigative Dermatology, 2014, 134, 712−718.
[5] B. Školová [Skolova], K. Hudská, P. Pullmannová, A. Kováčik, K. Palát, J. Roh, J. Fleddermann, I. Estrela-Lopis, K. Vávrová. Journal of Physical Chemistry B, 2014, 118, 10460 doi:10.1021/jp506407r
[6] I. Iwai, H. M. Han, L. den Hollander, S. Svensson, L. G. Öfverstedt, J. Anwar, J. Brewer, M. Bloksgaard, A. Laloeuf, D. Nosek, S. Masich, L. A. Bagatolli, U. Skoglund, Lars Norlén. Journal of Investigative Dermatology, 2012, 132, 2215–2225;. doi:10.1038/jid.2012.43.