Lipids in the Cell Cycle
The question of where we have come from has fascinated—and frustrated—intellectuals of all sorts, for centuries. There have therefore been experiments in several learned disciplines to investigate this problem. Between them, they have produced lots of answers to the question. One such discipline is cell biology. The question in this context is how a newly born human baby, a viable multi-cellular organism, can form from just one cell. A human live birth at full term comprises about 716 billion cells. This means that cell division produces an average of 2.6 billion cells per day of the pregnancy. It is hard to imagine that—but the number itslef is not that important: all we need to know for these purposes is that it is a lot.
Unfortunately, there are analogous and far less happy examples of cellular growth at this speed. Cancers typically start with a mutation in a single cell, forming one that typically divides so rapidly and without a means to stop it, that it can overwhelm the system in which it lives. The system of controlling cell division, and thus the cell cycle, has got lost, been turned off or been inhibited out of use. Both systems containing cancers, and healthy ones that do not have been researched by cell biologists, with a view to determining how their cell cycles are controlled.
The work of Sir Paul Nurse and Sir Tim Hunt (described on the and in a BBC documentary established the way the cell cycle is controlled in eukaryotes. It is controlled by changes in gene expression, by changes in the concentration of proteins called cyclins.
I first heard about this in 2010, when I was writing my PhD thesis. This was years after the Nobel prize has been awarded in 2001 and therefore longer still after it had been published. This sat in my mind until about two years ago, when I was researching E. coli for a quite different purpose. I stumbled upon evidence that there are no analogues of cyclins in bacteria, and that gene expression does not change in bacteria during the cell cycle [1]. What interested me about this work was that the question of what was controlling the cell cycle.
I looked into other details of the system. For example, structural proteins are made and assemble into a sort of internal scaffold that supports the physical process of division. However, their concentration does not change through the cell cycle, so it is not reasonable to think they are at the centre of controlling the cycle either [2].
One possibility, I thought, was a fundamental of the process. In order for a cell to divide, it must grow longer (elongate) in order that the two cells produced are large enough to be viable. This means that the cell envelope—the system comprising the cell membrane—must increase in size by a factor of two before division. So, at the very least, the membrane limits the rate of cell division. I developed the hypothesis that it controls it, too.
The principal component of the membrane is the lipid fraction. There are proteins as well, some of which produce lipids, but ultimately most of the membrane’s area is lipids. This means that in order for the cell to divide, a big effort in making lipids is required. This is simplified in E. coli because it comprises only about 4 major lipid components. One of these, phosphatidic acid (PA), typically represents less than 1% of the total and is thus often ignored. Another component, phosphatidylethanolamine (PE) often represents more than 80% of the total and thus dominates. Of the two remaining, phosphatidyl glycerol (PG) is the principal substrate for making cardolipin (CL).
Furthermore, evidence about how these lipids are spread out through the membrane, has been researched and reported. There is evidence from studies where cells were dyed that CL is found at the ends (poles) of the cells. This meant that when cells were elongated, the percentage of CL would be lower. This is because the poles have stayed at the same size but the middle of the cell is longer. This offered the tantalising hypothesis that the cell might control its lipid profile, only producing CL when it made new poles, for example during division.
We therefore tested the hypothesis that the lipid profile of E. coli changes through the cell cycle. The next obvious question to answer, if such a study was to take place, was how this should be done. We elected to arrest the cells in two of the three periods of the cell cycle, the resting state and the state in which they are at their most elongated. The lipid profile was determined at each (See figure). The results were that the CL fraction fell in the way we expected, but also that the PG fraction increased a lot. In fact, there was a higher percentage of PG in the elongated cells than in the ones in the resting state. This suggested that the rate of its production was faster than that of the cell’s elongation [3].
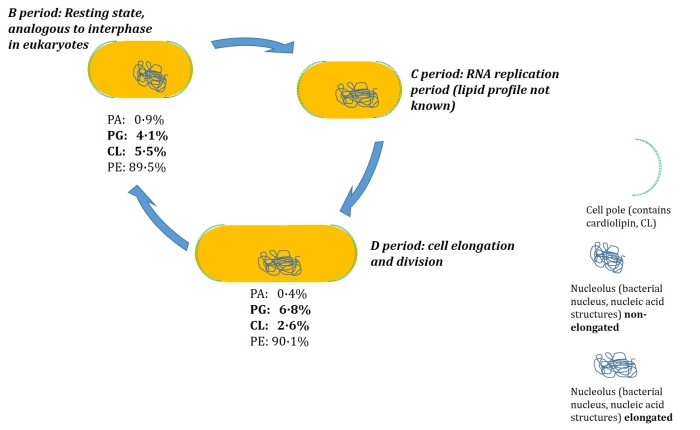
Figure. The cell division cycle in the bacteriumE. coli MG1655. The resting state (B period) of the cell is the most familiar, and in cell division is the one that progresses to replication of the nucleic acid structures (nucleolus, C period) before full elongation in the D period, during which the cell undergoes physical division. The representation of the D period is the state of the cell when arrested by rifampicin: the cell has elongated fully, but the system is unable to move closer to division. Lipid profiling data from reference [3].
This fast production of PG is consistent with the considerable evidence that it is the principal substrate for the synthesis of CL in vivo. The last result was that the PA fraction also appears to fall when the cells elongate, suggesting that it too is found at the poles, with CL. These results led us to conclude that lipid synthesis is controlled as a function of the cell cycle, with synthesis of the different lipids turned on and off [3].
These results raise several questions. One is whether this type of controlled lipid production is the norm in bacteria. Work by Yao et al. [4], also published this month, suggests that another Gram-negative bacterium, Chlamydia trachomatis has a very different way of making its membrane than previously thought. The reduced genome of this organism had led scientists to the conclusion that it acquired lipids from its host cells. However, this recent work shows that C. trachomatis has an approach to lipid production that appears to be quite similar to that established for E. coli. This work provides compelling evidence that this bacterium can produce its own PG, CL and PE from PA, rather than acquiring its lipids from the host as previously thought. Not only does this insight inform us about the control this bacterium has over its membrane, but it suggests that lipid production may be similar across a variety of Gram-negative bacteria.
Another question that the work on the lipid profile of E. coli as a function of the cell cycle raises, is whether anything similar happens in quite different cell types, such as human cells. Atilla-Gokcumen et al. [5] and Hague et al. [6] who showed, by arresting HeLa cells at slightly different points of the cell cycle, that the lipid profile of the membrane of this cell type changes as a function of the cell cycle. Atilla-Gokcumen et al. went further still, showing that lipid distribution and the physical properties of the lipid mixtures also changes through the cell cycle [5].
It is not clear what impact the change to the lipid profile in HeLa cell membranes has on the control mechanism of the cell cycle in these cells. The control mechanism I mentioned towards the beginning, that involves cyclins, makes it too difficult to be clear about this mechanism from the current data alone. However, it raises an interesting possible comparison. On one hand, there are eukaryotes who have developed from prokaryotes, and comprise a protein/gene-expression based cell cycle control system but in at least one cell type have significant change in lipid profile through the cycle. On the other, prokaryotes do not have the same protein-control system for the cell cycle as eukaryotes do, but also change their lipid composition through the cell cycle. Is change to the lipid profile through the cell cycle thus a fundamental part of cell division, whether or not it is the controlling mechanism?
There is much work to do to establish this, and so the story stops here for now. However, it may well go further forward in the next few years as it is conceivable that answering this question could support other human endeavour. This gives a social argument for funding. For those interested in the social and human angles of scientific research, the applications of understanding how the cell cycle of bacteria is controlled are at least two-fold. In cases where we might wish to stop them growing (in infection control) or to promote it (in industrial fermentation), what limits the cell cycle is important, as it provides both what can be exploited to arrest growth, and what must be supported to promote it.
References
[1] S. J. R. Arends, D. S. Weiss, Journal of Bacteriology, 2004, 186, 880-884. DOI: 10.1128/JB.186.3.880-884.2004
[2] S. Rueda, M. Vicente, J. Mingorance, Journal of Bacteriology, 2003, 185, 3344-3351. DOI: 10.1128/JB.185.11.3344-3351.2003
[3] S. Furse, H. Wienk, R. Boelens, A. I. P. M. de Kroon, J. A. Killian, FEBS Lett., 2015. DOI: 10.1016/j.febslet.2015.07.043
[4] J. Yao, P. T. Cherian, M. W. Frank, C. O. Rock, Journal of Biological Chemistry, 2015, 290, 18874 –18888. DOI: 10.1074/jbc.M115.657148
[5] G. E. Atilla-Gokcumen, E. Muro, J. Relat-Goberna, S. Sasse, A. Bedigian, M. L. Coughlin, S. Garcia-Manyes, U. S. Eggert, Cell, 2014, http://dx.doi.org/10.1016/j.cell.2013.12.015.
[6] C. V. Hague, A. D. Postle, G. S. Attard, M. K. Dymond, Faraday Discussions 2013 161, 481-497. DOI: http://dx.doi.org/10.1039/c2fd20078c
Conflict of interest statement: I am the first author of reference [3].