A Patch on the Membrane
Sickle-cell anaemia and brittle bone disease are both caused by a single, small change in the structure of a protein. Cystic fibrosis is also caused by one of several ‘point’ mutations in the structure of just one protein. The question a scientist might ask is why this should occur—why should a small change in structure result in such a striking change in behaviour?
A practical difficulty in answering this question is how to test for the functional difference between normal and disease mutant forms of a protein. Cytosolic proteins can be characterised in aqueous systems that are effectively a model of the liquid part of cells. This means that the protein that is at fault in sickle cell anaemiacs can be characterised reasonably easily. The functional behaviour of the protein in cystic fibrosis is more difficult to characterise because it is a membrane protein rather than a cytosolic one.
Like lipids, membrane proteins have a part that likes water (hydrophillic) and one that does not (hydrophobic). This means that on contact with water without a membrane, they can re-fold, i.e. morph into a bit of a mess. The difficulty with a lot of membrane proteins—both disease and normal types—is that isolating them so that they can be characterised is an insurmountable obstacle.
The practical difficulty is therefore to isolate the protein in its native protein conformation. Detergents have been used for this purpose, and these do go some way to satisfying the amphiphilic character of membrane proteins. However, they can twist them in other conformers, and that can be just a difficult to understand, and just as useless in researching a disease. A better idea is surely to isolate the proteins with a patch of membrane that they like, such as the one they have naturally?
This is exactly what several groups of scientists have done [1-3]. The heart of this technique is to use a sort of molecular belt that acts like a polymerised lipid, with the greasy part of the lipid facing the fatty acid residues of the lipids and the hydrophilic part of the polymer facing the aqueous fraction. It turns out that this is thermodynamically stable and that sections of this polymerised lipid of the right length can behave like pastry cutters and chop out little discs of the membrane, with the all-important membrane protein nonchalantly sitting in the middle, like a prince in a sedan chair (Figure).
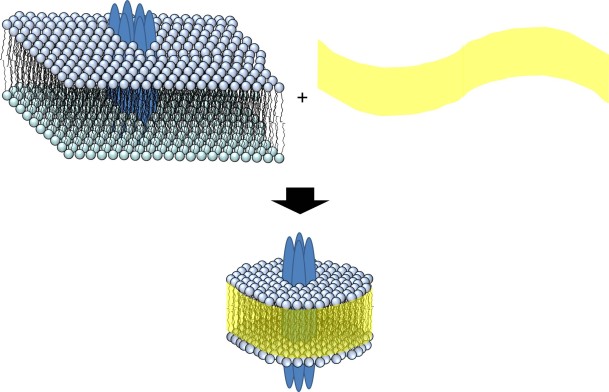
Figure. A section of the membrane is separated from the rest by the addition of co-polymer (yellow) to give a nanodisc (bottom).
Long et al. [1] showed that several mitochondrial proteins could be isolated this way. They established this by showing that the proteins isolated still had their enzyme activity after the isolation. This provided excellent data that functioning proteins resident in the membrane can be isolated for studies conducted in vitro.
Swainsbury et al. [2] tested different membrane protein isolation methods using a photoreaction centre from a purple bacterium called Rhodobacter sphaeroides. This protein complex is a useful tool for comparing methods because its behaviour with respect to light is dependent upon its conformation. In fact, both small and large differences in the conformation of the protein can be measured.
Doerr et al. [3] chose a different and more difficult target, an ion channel. Ion channels transport metal ions like sodium and potassium across membranes. The difficulty in isolating and characterising these is that unless they are in a membrane that divides two aqueous compartments, the changes in concentration of the ions cannot be measured. It is the molecular equivalent of letting someone in by opening the door of a room that has a wall missing. What this group therefore did was to isolate the ion channel protein complex (called KcsA) and reconstitute it in a membrane that did indeed divide two aqueous compartments. Electrophysiology techniques were then used to establish the activity of the KcsA they isolated. They found that indeed it was active after being both put into these little membrane patches (nanodiscs) and then reconstituted into another membrane.
What this research shows is that a variety of proteins, with different functions and from different species, can be isolated and characterised using this approach. The work is at an early stage, meaning that there is the unfortunate but predictable rush to secure the Intellectual Property rights. One form of this is a registered trade mark that has suffered from 90s-style marketing, having the name ‘lipodisqs®’. This has also meant that the material for making nanodiscs has become commercially available. It is at a high price, the equivalent of £6m/Kg in the Netherlands.
However, as the facility of this technique becomes clear and more widely used, I predict that it will become easier to use and more finely tuned. We may yet see the characterisation of membrane proteins that exist in tiny quantities, such as those in individual neurone synapses. It may allow us to determine shifts in membrane protein concentration at different points in the cell division cycle. This technique may have scope to allow us to chart the shifts in molecular profile that represent the processes of a normal working body. However, its strongest application in the current zeitgeist may be to determine the functional difference between individual membrane proteins in diseased and healthy systems, enabling the characterisation of disease in a clearer and quicker way than ever before.
References
[1] A. R. Long, C. C. O’Brien, K. Malhotra, C. T. Schwall, A. D. Albert, A. Watts, N. N. Alder, BMC Biotechnology, 2013, 13, 41. DOI: 10.1186/1472-6750-13-41
[2] D. J. K. Swainsbury, S. Scheidelaar, R. van Grondelle, J. A. Killian, M. R. Jones, Angew. Chem. Int. Ed., 2014, 53, 11803. DOI: 10.1002/anie.201406412
[3] J. M. Doerr, M. C. Koorengevel, M. Schäfer, A. V. Prokofyev, S. Scheidelaar, E. A. W. van der Cruijsen, T. R. Dafforn, Marc Baldus, and J. A. Killian, Proc. Natl. Acad. Sci., 2014, DOI 10.1073/pnas.1416205112.
Conflict of Interest Statement: I share an office with the first author of [3]. The last author of the same paper is my boss, though I work on a separate project.